Publications
Publications in peer reviewed journals
Microclimate shapes the phylosymbiosis of rodent gut microbiota in Jordan's Great Rift Valley.
2023 - Front Microbiol, 1258775
Abstract:
Host phylogeny and the environment play vital roles in shaping animal microbiomes. However, the effects of these variables on the diversity and richness of the gut microbiome in different bioclimatic zones remain underexplored. In this study, we investigated the effects of host phylogeny and bioclimatic zone on the diversity and composition of the gut microbiota of two heterospecific rodent species, the spiny mouse and the house mouse , in three bioclimatic zones of the African Great Rift Valley (GRV). We confirmed host phylogeny using the sequencing method and analyzed the influence of host phylogeny and bioclimatic zone parameters on the rodent gut microbiome using high-throughput amplicon sequencing of 16S rRNA gene fragments. Phylogenetic analysis supported the morphological identification of the rodents and revealed a marked genetic difference between the two heterospecific species. We found that bioclimatic zone had a significant effect on the gut microbiota composition while host phylogeny did not. Microbial alpha diversity of heterospecific hosts was highest in the Mediterranean forest bioclimatic zone, followed by the Irano-Turanian shrubland, and was lowest in the Sudanian savanna tropical zone. The beta diversity of the two rodent species showed significant differences across the Mediterranean, Irano-Turanian, and Sudanian regions. The phyla and were highly abundant, and and were also prominent. Amplicon sequence variants (ASVs) were identified that were unique to the Sudanian bioclimatic zone. The core microbiota families recovered in this study were consistent among heterospecific hosts. However, diversity decreased in conspecific host populations found at lower altitudes in Sudanian bioclimatic zone. The composition of the gut microbiota is linked to the adaptation of the host to its environment, and this study underscores the importance of incorporating climatic factors such as elevation and ambient temperature, in empirical microbiome research and is the first to describe the rodent gut microbiome from the GRV.
Ecophysiology and interactions of a taurine-respiring bacterium in the mouse gut.
2023 - Nat Commun, 1: 5533
Abstract:
Taurine-respiring gut bacteria produce HS with ambivalent impact on host health. We report the isolation and ecophysiological characterization of a taurine-respiring mouse gut bacterium. Taurinivorans muris strain LT0009 represents a new widespread species that differs from the human gut sulfidogen Bilophila wadsworthia in its sulfur metabolism pathways and host distribution. T. muris specializes in taurine respiration in vivo, seemingly unaffected by mouse diet and genotype, but is dependent on other bacteria for release of taurine from bile acids. Colonization of T. muris in gnotobiotic mice increased deconjugation of taurine-conjugated bile acids and transcriptional activity of a sulfur metabolism gene-encoding prophage in other commensals, and slightly decreased the abundance of Salmonella enterica, which showed reduced expression of galactonate catabolism genes. Re-analysis of metagenome data from a previous study further suggested that T. muris can contribute to protection against pathogens by the commensal mouse gut microbiota. Together, we show the realized physiological niche of a key murine gut sulfidogen and its interactions with selected gut microbiota members.
Neutral and Pectic Heteropolysaccharides Isolated from Mucilage: Composition, Molecular Dimensions and Prebiotic Potential.
2023 - Int J Mol Sci, 4: in press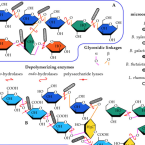
Abstract:
is a semi-wild cactus cultivated for its fruit. However, the cladodes are often discarded, wasting the potentially useful mucilage in them. The mucilage is composed primarily of heteropolysaccharides, characterized by their molar mass distribution, monosaccharide composition, structural features (by vibrational spectroscopy, FT IR, and atomic force microscopy, AFM), and fermentability by known saccharolytic commensal members of the gut microbiota. After fractionation with ion exchange chromatography, four polysaccharides were found: one neutral (composed mainly of galactose, arabinose, and xylose) and three acidic, with a galacturonic acid content from 10 to 35%. Their average molar masses ranged from 1.8 × 10 to 2.8 × 10 g·mol. Distinct structural features such as galactan, arabinan, xylan, and galacturonan motifs were present in the FT IR spectra. The intra- and intermolecular interactions of the polysaccharides, and their effect on the aggregation behavior, were shown by AFM. The composition and structural features of these polysaccharides were reflected in their prebiotic potential. and were not able to utilize them, whereas members of showed utilization capacity. The obtained data suggest a high economic potential for this species, with potential uses such as animal feed in arid areas, precise prebiotic, and symbiotic formulations, or as the carbon skeleton source in a green refinery. Our methodology can be used to evaluate the saccharides as the phenotype of interest, helping to guide the breeding strategy.
Gut microbiome signatures of Yorkshire Terrier enteropathy during disease and remission.
2023 - Sci Rep, 1: 4337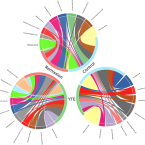
Abstract:
The role of the gut microbiome in developing Inflammatory Bowel Disease (IBD) in humans and dogs has received attention in recent years. Evidence suggests that IBD is associated with alterations in gut microbial composition, but further research is needed in veterinary medicine. The impact of IBD treatment on the gut microbiome needs to be better understood, especially in a breed-specific form of IBD in Yorkshire Terriers known as Yorkshire Terrier Enteropathy (YTE). This study aimed to investigate the difference in gut microbiome composition between YTE dogs during disease and remission and healthy Yorkshire Terriers. Our results showed a significant increase in specific taxa such as Clostridium sensu stricto 1, Escherichia-Shigella, and Streptococcus, and a decrease in Bacteroides, Prevotella, Alloprevotella, and Phascolarctobacterium in YTE dogs compared to healthy controls. No significant difference was found between the microbiome of dogs in remission and those with active disease, suggesting that the gut microbiome is affected beyond clinical recovery.
Differential carbon utilization enables co-existence of recently speciated Campylobacteraceae in the cow rumen epithelial microbiome.
2023 - Nat Microbiol, in press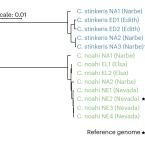
Abstract:
The activities of different microbes in the cow rumen have been shown to modulate the host's ability to utilize plant biomass, while the host-rumen interface has received little attention. As datasets collected worldwide have pointed to Campylobacteraceae as particularly abundant members of the rumen epithelial microbiome, we targeted this group in a subset of seven cows with meta- and isolate genome analysis. We show that the dominant Campylobacteraceae lineage has recently speciated into two populations that were structured by genome-wide selective sweeps followed by population-specific gene import and recombination. These processes led to differences in gene expression and enzyme domain composition that correspond to the ability to utilize acetate, the main carbon source for the host, at the cost of inhibition by propionate. This trade-off in competitive ability further manifests itself in differential dynamics of the two populations in vivo. By exploring population-level adaptations that otherwise remain cryptic in culture-independent analyses, our results highlight how recent evolutionary dynamics can shape key functional roles in the rumen microbiome.
Pathometagenomics reveals susceptibility to intestinal infection by Morganella to be mediated by the blood group-related B4galnt2 gene in wild mice.
2023 - Gut Microbes, 1: 2164448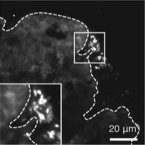
Abstract:
Infectious disease is widely considered to be a major driver of evolution. A preponderance of signatures of balancing selection at blood group-related genes is thought to be driven by inherent trade-offs in susceptibility to disease. B4galnt2 is subject to long-term balancing selection in house mice, where two divergent allele classes direct alternative tissue-specific expression of a glycosyltransferase in the intestine versus blood vessels. The blood vessel allele class leads to prolonged bleeding times similar to von Willebrand disease in humans, yet has been maintained for millions of years. Based on in vivo functional studies in inbred lab strains, it is hypothesized that the cost of prolonged bleeding times may be offset by an evolutionary trade-off involving susceptibility to a yet unknown pathogen(s). To identify candidate pathogens for which resistance could be mediated by B4galnt2 genotype, we here employed a novel "pathometagenomic" approach in a wild mouse population, which combines bacterial 16S rRNA gene-based community profiling with histopathology of gut tissue. Through subsequent isolation, genome sequencing and controlled experiments in lab mice, we show that the presence of the blood vessel allele is associated with resistance to a newly identified subspecies of Morganella morganii, a clinically important opportunistic pathogen. Given the increasing importance of zoonotic events, the approach outlined here may find useful application in the detection of emerging diseases in wild animal populations.